بازدید:34827 بروزرسانی: 20-12-1402
Click here or contact me to download PARS software.
PARS is designed for calculating the number of sequence fragments (mono, di and three-peptides), with assigning their conformation in secondary structure terms. As input, you need the structural file format, *.DSSP, for your structure. This file format is a plain text file format with DSSP extension, containing necessary secondary structure data for any residues.
Working with PDB structural file format (obtained from Protein Data Bank), you have to convert it to DSSP format. This can be done in some ways. See these pages for more information:
http://www.cmbi.ru.nl/dssp.html
or
A DSSP file format sample is here.
After preparing your input, you can use PARS software. Here is graphical step by step help:
(Crystal Structure of Bovine Serum Albumin, PDB ID: 4f5s explained here)
1
- Load your input:
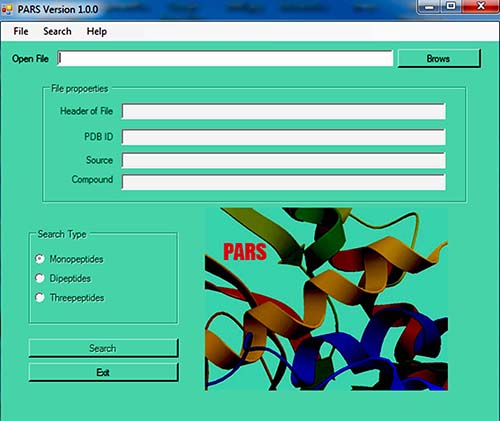
Calculate fragment frequencies and save the results:
A sample of graphical output:
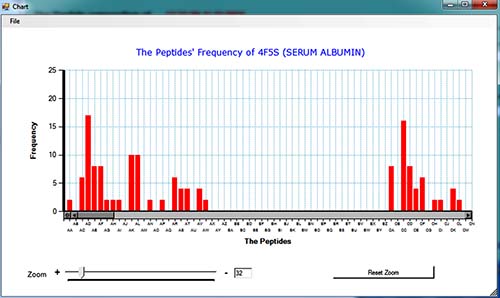
Also, the outputs are savable as two format, both in PARS extensions. (Format1, Format2)
For any question, specially reporting bugs, contact: heshmati@znu.ac.ir or bioinf.comp.biol@gmail.com

